Cell Biology, Molecular Biology, and Genetics - MCAT Biological and Biochemical Foundations of Living Systems
Card 1 of 2944
Sexually transmitted diseases are a common problem among young people in the United States. One of the more common diseases is caused by the bacterium Neisseria gonorrhoeae, which leads to inflammation and purulent discharge in the male and female reproductive tracts.
The bacterium has a number of systems to evade host defenses. Upon infection, it uses pili to adhere to host epithelium. The bacterium also uses an enzyme, gonococcal sialyltransferase, to transfer a sialyic acid residue to a gonococcal surface lipooligosaccharide (LOS). A depiction of this can be seen in Figure 1. The sialyic acid residue mimics the protective capsule found on other bacterial species.
Once infection is established, Neisseria preferentially infects columnar epithelial cells in the female reproductive tract, and leads to a loss of cilia on these cells. Damage to the reproductive tract can result in pelvic inflammatory disease, which can complicate pregnancies later in the life of the woman.

A scientist discovers that Neisseria uses flagella to move between host cells. Which of the following is true of the flagella?
Sexually transmitted diseases are a common problem among young people in the United States. One of the more common diseases is caused by the bacterium Neisseria gonorrhoeae, which leads to inflammation and purulent discharge in the male and female reproductive tracts.
The bacterium has a number of systems to evade host defenses. Upon infection, it uses pili to adhere to host epithelium. The bacterium also uses an enzyme, gonococcal sialyltransferase, to transfer a sialyic acid residue to a gonococcal surface lipooligosaccharide (LOS). A depiction of this can be seen in Figure 1. The sialyic acid residue mimics the protective capsule found on other bacterial species.
Once infection is established, Neisseria preferentially infects columnar epithelial cells in the female reproductive tract, and leads to a loss of cilia on these cells. Damage to the reproductive tract can result in pelvic inflammatory disease, which can complicate pregnancies later in the life of the woman.

A scientist discovers that Neisseria uses flagella to move between host cells. Which of the following is true of the flagella?
Tap to reveal answer
Flagella and cilia share a 9+2 ultrastructure, where there is a ring of 9 doublets of microtubules surrounding a central doublet, and connects via dynein arms.
Flagella and cilia share a 9+2 ultrastructure, where there is a ring of 9 doublets of microtubules surrounding a central doublet, and connects via dynein arms.
← Didn't Know|Knew It →
Choose the transcript created if RNA polymerase transcribes the following template strand.
5'-ACTTGCAGGCC-3'
Choose the transcript created if RNA polymerase transcribes the following template strand.
5'-ACTTGCAGGCC-3'
Tap to reveal answer
When transcribing from a template strand, here are a few things to remember:
1. RNA polymerase reads the strand in the 3' to 5' direction.
2. In the RNA transcript, thymine is replaced with uracil.
In order to double check, make sure that the two strands are complementary when antiparallel to one another.
Template: 5'-ACTTGCAGGCC-3'
Transcript: 3'-UGAACGUCCGG-5'
When transcribing from a template strand, here are a few things to remember:
1. RNA polymerase reads the strand in the 3' to 5' direction.
2. In the RNA transcript, thymine is replaced with uracil.
In order to double check, make sure that the two strands are complementary when antiparallel to one another.
Template: 5'-ACTTGCAGGCC-3'
Transcript: 3'-UGAACGUCCGG-5'
← Didn't Know|Knew It →
Which of the following organelles is most important in testosterone synthesis?
Which of the following organelles is most important in testosterone synthesis?
Tap to reveal answer
Testosterone is a steroid hormone. Steroids are nonpolar hormones; they start off as cholesterol and are converted to steroids in the cytosol. The cholesterol necessary to create steroids are synthesized in the smooth endoplasmic reticulum. Crucial functions of the smooth endoplasmic reticulum include lipid synthesis and detoxification.
Ribosomes are responsible for protein synthesis in translation and help generate numerous peptide hormones. The rough endoplasmic reticulum helps with vesicle formation and transport. The nucleolus generates the biological materials to build ribosomes.
Testosterone is a steroid hormone. Steroids are nonpolar hormones; they start off as cholesterol and are converted to steroids in the cytosol. The cholesterol necessary to create steroids are synthesized in the smooth endoplasmic reticulum. Crucial functions of the smooth endoplasmic reticulum include lipid synthesis and detoxification.
Ribosomes are responsible for protein synthesis in translation and help generate numerous peptide hormones. The rough endoplasmic reticulum helps with vesicle formation and transport. The nucleolus generates the biological materials to build ribosomes.
← Didn't Know|Knew It →
Which of the following proteins would be transcribed at the level of the rough endoplasmic reticulum?
Which of the following proteins would be transcribed at the level of the rough endoplasmic reticulum?
Tap to reveal answer
It is important for students to understand that transmembrane proteins, as well as secreted proteins, need to be synthesized at the level of the rough endoplasmic reticulum. All of the other options are examples of proteins that exert their function in the cytosol, and thus are transcribed by free ribosomes in the cytosol.
It is important for students to understand that transmembrane proteins, as well as secreted proteins, need to be synthesized at the level of the rough endoplasmic reticulum. All of the other options are examples of proteins that exert their function in the cytosol, and thus are transcribed by free ribosomes in the cytosol.
← Didn't Know|Knew It →
What is the difference between cytosolic ribosomes and ribosomes on the rough endoplasmic reticulum (RER)?
What is the difference between cytosolic ribosomes and ribosomes on the rough endoplasmic reticulum (RER)?
Tap to reveal answer
While both types of ribosomes are used to make proteins, the difference between them has to do with the fate of the proteins. Cytosolic ribosomes make proteins for the cytosol, which rough endoplasmic reticulum ribosomes make them to be bound in membranes, or to be excreted from the cell in vesicles (exocytosis). Both types of ribosmes require a peptidyl transferase to elongate the peptide chain during protein synthesis.
While both types of ribosomes are used to make proteins, the difference between them has to do with the fate of the proteins. Cytosolic ribosomes make proteins for the cytosol, which rough endoplasmic reticulum ribosomes make them to be bound in membranes, or to be excreted from the cell in vesicles (exocytosis). Both types of ribosmes require a peptidyl transferase to elongate the peptide chain during protein synthesis.
← Didn't Know|Knew It →
Scientists identify a mutation in an isolated community in central Africa that prevents individuals from detoxifying potentially harmful organic molecules, leading to a high percentage of people who become very ill after consuming alcohol. What cellular organelle does this mutation most likely affect the most?
Scientists identify a mutation in an isolated community in central Africa that prevents individuals from detoxifying potentially harmful organic molecules, leading to a high percentage of people who become very ill after consuming alcohol. What cellular organelle does this mutation most likely affect the most?
Tap to reveal answer
The smooth endoplasmic reticulum functions in drug and alcohol detoxification. A common incorrect answer chosen here is the lysozome, which digests food, bacteria, viruses, and damaged organelles or cellular structures.
The smooth endoplasmic reticulum functions in drug and alcohol detoxification. A common incorrect answer chosen here is the lysozome, which digests food, bacteria, viruses, and damaged organelles or cellular structures.
← Didn't Know|Knew It →
Scientists use a process called Flourescent In-Situ Hybridization, or FISH, to study genetic disorders in humans. FISH is a technique that uses spectrographic analysis to determine the presence or absence, as well as the relative abundance, of genetic material in human cells.
To use FISH, scientists apply fluorescently-labeled bits of DNA of a known color, called probes, to samples of test DNA. These probes anneal to the sample DNA, and scientists can read the colors that result using laboratory equipment. One common use of FISH is to determine the presence of extra DNA in conditions of aneuploidy, a state in which a human cell has an abnormal number of chromosomes. Chromosomes are collections of DNA, the totality of which makes up a cell’s genome. Another typical use is in the study of cancer cells, where scientists use FISH labels to ascertain if genes have moved inappropriately in a cell’s genome.
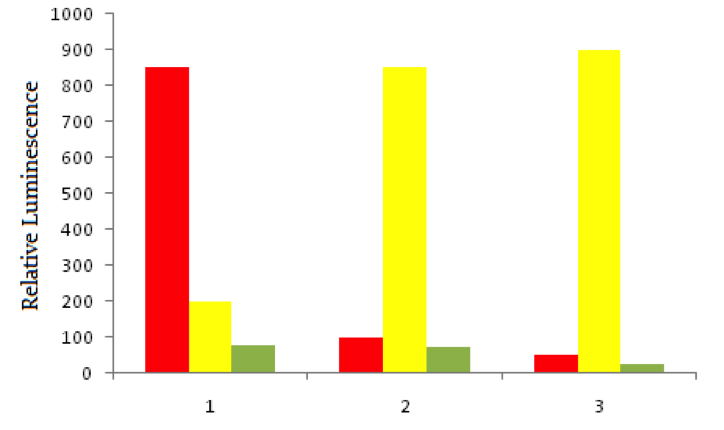
Using red fluorescent tags, scientists label probe DNA for a gene known to be expressed more heavily in cancer cells than normal cells. They then label a probe for an immediately adjacent DNA sequence with a green fluorescent tag. Both probes are then added to three dishes, shown below. In dish 1 human bladder cells are incubated with the probes, in dish 2 human epithelial cells are incubated, and in dish 3 known non-cancerous cells are used. The relative luminescence observed in regions of interest in all dishes is shown below.
A scientist discovers that there is a class of proteins called tumor suppressors. These proteins are present in the cytosol of almost all human cells, and serve to downregulate cell division by preventing entry into key parts of the cell cycle. Where are these proteins most likely synthesized?
Scientists use a process called Flourescent In-Situ Hybridization, or FISH, to study genetic disorders in humans. FISH is a technique that uses spectrographic analysis to determine the presence or absence, as well as the relative abundance, of genetic material in human cells.
To use FISH, scientists apply fluorescently-labeled bits of DNA of a known color, called probes, to samples of test DNA. These probes anneal to the sample DNA, and scientists can read the colors that result using laboratory equipment. One common use of FISH is to determine the presence of extra DNA in conditions of aneuploidy, a state in which a human cell has an abnormal number of chromosomes. Chromosomes are collections of DNA, the totality of which makes up a cell’s genome. Another typical use is in the study of cancer cells, where scientists use FISH labels to ascertain if genes have moved inappropriately in a cell’s genome.
Using red fluorescent tags, scientists label probe DNA for a gene known to be expressed more heavily in cancer cells than normal cells. They then label a probe for an immediately adjacent DNA sequence with a green fluorescent tag. Both probes are then added to three dishes, shown below. In dish 1 human bladder cells are incubated with the probes, in dish 2 human epithelial cells are incubated, and in dish 3 known non-cancerous cells are used. The relative luminescence observed in regions of interest in all dishes is shown below.

A scientist discovers that there is a class of proteins called tumor suppressors. These proteins are present in the cytosol of almost all human cells, and serve to downregulate cell division by preventing entry into key parts of the cell cycle. Where are these proteins most likely synthesized?
Tap to reveal answer
Tumor suppressors, as defined in the question, are proteins. Proteins are synthesized by ribosomes. While the rough endoplasmic reticulum contains ribosomes, its function is related to the synthesis of transmembrane and extra-cellular proteins. Cytosolic ribosomes will be responsible for synthesis of proteins that remain in the cell.
Tumor suppressors, as defined in the question, are proteins. Proteins are synthesized by ribosomes. While the rough endoplasmic reticulum contains ribosomes, its function is related to the synthesis of transmembrane and extra-cellular proteins. Cytosolic ribosomes will be responsible for synthesis of proteins that remain in the cell.
← Didn't Know|Knew It →
Which of the following cellular components synthesizes lipids of the plasma membrane?
Which of the following cellular components synthesizes lipids of the plasma membrane?
Tap to reveal answer
Lipids that are usually used for the cell membrane are created in the smooth endoplasmic reticulum. The rough endoplasmic reticulum and ribosomes are involved in protein production. The mitochondria are essential in producing ATP.
Lipids that are usually used for the cell membrane are created in the smooth endoplasmic reticulum. The rough endoplasmic reticulum and ribosomes are involved in protein production. The mitochondria are essential in producing ATP.
← Didn't Know|Knew It →
Where is the eukaryotic electron transport chain located?
Where is the eukaryotic electron transport chain located?
Tap to reveal answer
The electron transport chain (ETC) is located on the inner mitochondrial membrane. It is composed of a set of cytochromes which function in creating a proton (H+) gradient which provides the energy for ATP synthase to make ATP.
Note that the prokaryotic electron transport chain occurs on the plasma membrane with a proton gradient generated between two outer membranes or between the membrane and cell wall.
The electron transport chain (ETC) is located on the inner mitochondrial membrane. It is composed of a set of cytochromes which function in creating a proton (H+) gradient which provides the energy for ATP synthase to make ATP.
Note that the prokaryotic electron transport chain occurs on the plasma membrane with a proton gradient generated between two outer membranes or between the membrane and cell wall.
← Didn't Know|Knew It →
In 2013, scientists linked a cellular response called the unfolded protein response (UPR) to a series of neurodegenerative diseases, including such major health issues as Parkinson’s and Alzheimer’s Disease. According to their work, the unfolded protein response is a reduction in translation as a result of a series of enzymes that modify a translation initiation factor, eIF2, as below:

In the above sequence, the unfolded protein sensor binds to unfolded protein, such as the pathogenic amyloid-beta found in the brains of Alzheimer’s Disease patients. This sensor then phosphorylates PERK, or protein kinase RNA-like endoplasmic reticulum kinase. This leads to downstream effects on eIF2, inhibition of which represses translation. It is thought that symptoms of neurodegenerative disease may be a result of this reduced translation.
Regarding unfolded proteins discussed in the passage, which organelle is likely to be the site of initial protein folding in normal cells?
In 2013, scientists linked a cellular response called the unfolded protein response (UPR) to a series of neurodegenerative diseases, including such major health issues as Parkinson’s and Alzheimer’s Disease. According to their work, the unfolded protein response is a reduction in translation as a result of a series of enzymes that modify a translation initiation factor, eIF2, as below:

In the above sequence, the unfolded protein sensor binds to unfolded protein, such as the pathogenic amyloid-beta found in the brains of Alzheimer’s Disease patients. This sensor then phosphorylates PERK, or protein kinase RNA-like endoplasmic reticulum kinase. This leads to downstream effects on eIF2, inhibition of which represses translation. It is thought that symptoms of neurodegenerative disease may be a result of this reduced translation.
Regarding unfolded proteins discussed in the passage, which organelle is likely to be the site of initial protein folding in normal cells?
Tap to reveal answer
Protein folding takes place in the endoplasmic reticulum, typically coinciding with the translation by bound ribosomes of the rough endoplasmic reticulum. Further modification can take place in the Golgi. Note that ribosomes in the cytosol or on the rough endoplasmic reticulum may translate a protein, but the protein folding will occur in the endoplasmic reticulum.
Protein folding takes place in the endoplasmic reticulum, typically coinciding with the translation by bound ribosomes of the rough endoplasmic reticulum. Further modification can take place in the Golgi. Note that ribosomes in the cytosol or on the rough endoplasmic reticulum may translate a protein, but the protein folding will occur in the endoplasmic reticulum.
← Didn't Know|Knew It →
The liver is one of the major sites for drug metabolism and detoxification. Which organelle would you expect to play an important role in this process?
The liver is one of the major sites for drug metabolism and detoxification. Which organelle would you expect to play an important role in this process?
Tap to reveal answer
One of the major functions of the smooth endoplasmic reticulum is drug metabolism. In the liver, the cellular smooth endoplasmic reticulum serves as the primary site of drug and alcohol detoxifiction. This organelle is present (and active) in cells throughout the body, but plays the most impactful role in the cells of the liver. While lysosomes are responsible for clearing cellular debris, they commonly digest pathogens and microbes rather than chemical contaminants, like drugs or alcohol.
One of the major functions of the smooth endoplasmic reticulum is drug metabolism. In the liver, the cellular smooth endoplasmic reticulum serves as the primary site of drug and alcohol detoxifiction. This organelle is present (and active) in cells throughout the body, but plays the most impactful role in the cells of the liver. While lysosomes are responsible for clearing cellular debris, they commonly digest pathogens and microbes rather than chemical contaminants, like drugs or alcohol.
← Didn't Know|Knew It →
Type 1 diabetes is a well-understood autoimmune disease. Autoimmune diseases result from an immune system-mediated attack on one’s own body tissues. In normal development, an organ called the thymus introduces immune cells to the body’s normal proteins. This process is called negative selection, as those immune cells that recognize normal proteins are deleted. If cells evade this process, those that recognize normal proteins enter into circulation, where they can attack body tissues. The thymus is also important for activating T-cells that recognize foreign proteins.
As the figure below shows, immune cells typically originate in the bone marrow. Some immune cells, called T-cells, then go to the thymus for negative selection. Those that survive negative selection, enter into general circulation to fight infection. Other cells, called B-cells, directly enter general circulation from the bone marrow. It is a breakdown in this carefully orchestrated process that leads to autoimmune disease, such as type 1 diabetes.

T-cells use receptors in their activity to defend their biological hosts. These receptors are protein molecules, heavily modified before being sent to the cell's surface. In which organelle does the majority of such modification take place?
Type 1 diabetes is a well-understood autoimmune disease. Autoimmune diseases result from an immune system-mediated attack on one’s own body tissues. In normal development, an organ called the thymus introduces immune cells to the body’s normal proteins. This process is called negative selection, as those immune cells that recognize normal proteins are deleted. If cells evade this process, those that recognize normal proteins enter into circulation, where they can attack body tissues. The thymus is also important for activating T-cells that recognize foreign proteins.
As the figure below shows, immune cells typically originate in the bone marrow. Some immune cells, called T-cells, then go to the thymus for negative selection. Those that survive negative selection, enter into general circulation to fight infection. Other cells, called B-cells, directly enter general circulation from the bone marrow. It is a breakdown in this carefully orchestrated process that leads to autoimmune disease, such as type 1 diabetes.

T-cells use receptors in their activity to defend their biological hosts. These receptors are protein molecules, heavily modified before being sent to the cell's surface. In which organelle does the majority of such modification take place?
Tap to reveal answer
The Golgi apparatus is the organelle tasked specifically with the modification of proteins. Membrane proteins are synthesized by ribosomes of the rough endoplasmic reticulum before being sent to the Golgi apparatus for modification and packaging. This packaging allows for integration of the receptor proteins into the membrane.
The Golgi apparatus is the organelle tasked specifically with the modification of proteins. Membrane proteins are synthesized by ribosomes of the rough endoplasmic reticulum before being sent to the Golgi apparatus for modification and packaging. This packaging allows for integration of the receptor proteins into the membrane.
← Didn't Know|Knew It →
In eukaryotic cells, what organelle is associated with translation of antibody proteins?
In eukaryotic cells, what organelle is associated with translation of antibody proteins?
Tap to reveal answer
Antibody proteins are translated by ribosomes attached to the rough endoplasmic reticulum, and subsequently secreted out of the cell in vesicles. While the Golgi apparatus is involved in the secretory pathway of these proteins, it is not specifically involved in the translation.
The nucleus and mitochondria are not associated with antibody protein translation.
Antibody proteins are translated by ribosomes attached to the rough endoplasmic reticulum, and subsequently secreted out of the cell in vesicles. While the Golgi apparatus is involved in the secretory pathway of these proteins, it is not specifically involved in the translation.
The nucleus and mitochondria are not associated with antibody protein translation.
← Didn't Know|Knew It →
A vesicle travelling from the back to the endoplasmic reticulum is most likely contains a protein coat.
A vesicle travelling from the back to the endoplasmic reticulum is most likely contains a protein coat.
Tap to reveal answer
There are three common vesicle protein coats: clathrin, COPI, and COPII. Vesicles coated in clathrin are typically being sent from the Golgi to the plasma membrane or endosomes. COPII vesicles are headed from the endoplasmic reticulum to the Golgi body. Vesicles coated in COPI are typically involved in retrograde transport from the Golgi body to the endoplasmic reticulum (recycling proteins).
There are three common vesicle protein coats: clathrin, COPI, and COPII. Vesicles coated in clathrin are typically being sent from the Golgi to the plasma membrane or endosomes. COPII vesicles are headed from the endoplasmic reticulum to the Golgi body. Vesicles coated in COPI are typically involved in retrograde transport from the Golgi body to the endoplasmic reticulum (recycling proteins).
← Didn't Know|Knew It →
Which of the following choices describe functions of the Golgi apparatus?
I. Post-translational modifications
II. Formation of lysosomes
III. Carbohydrate synthesis
IV. Protein trafficking
Which of the following choices describe functions of the Golgi apparatus?
I. Post-translational modifications
II. Formation of lysosomes
III. Carbohydrate synthesis
IV. Protein trafficking
Tap to reveal answer
Every choice describes one of the diverse functions of the Golgi apparatus. It is a crucial organelle for protein trafficking via the secretory pathway. It also serves as one of the first steps of lysosome formation, by organizing lysosomal proteins and making vesicles that will eventually become mature lysosomes. The Golgi is also responsible for various post-translational modifications, including processing of glycosylation. Carbohydrate synthesis also occurs in the Golgi, facilitating glycosylation and other modification pathways.
Every choice describes one of the diverse functions of the Golgi apparatus. It is a crucial organelle for protein trafficking via the secretory pathway. It also serves as one of the first steps of lysosome formation, by organizing lysosomal proteins and making vesicles that will eventually become mature lysosomes. The Golgi is also responsible for various post-translational modifications, including processing of glycosylation. Carbohydrate synthesis also occurs in the Golgi, facilitating glycosylation and other modification pathways.
← Didn't Know|Knew It →
There are two models for the operation of the Golgi apparatus in eukaryotic cells. As it is difficult to visualize the operation of cells at the molecular level in real time, scientists typically rely on static electron micrographs to see the morphology of organelles. As a result, the dynamic operation of these organelles can sometimes be unclear.
Cisternal Maturation Hypothesis
In the cisternal maturation hypothesis, the cisternae of the Golgi apparatus evolve. Proteins leave the endoplasmic reticulum, and enter the cis-Golgi. The cisterna of the cis-Golgi then matures, with its enzymatic contents and internal environment changing as it becomes the medial-Golgi, and, eventually, the trans-Golgi.
In this model, the proteins never physically leave their membrane-bound cisternae during their transit across the Golgi. Instead, the entire unit of contents remains within the evolving cisternae.
Vesicular Transport Hypothesis
In contrast to the cisternal maturation hypothesis, the vesicular transport hypothesis posits that the cis-, medial-, and trans-Golgi cisternae are more static structures. Instead of evolving around their contents, the contents are physically shuttled via vesicular intermediates from each cisterna to the next.
In the case of vesicular transport, vesicles are shuttled along microtubules. Motor proteins facilitate this movement, with unique proteins being used for each direction of movement along a microtubule.
Which of the following is most likely to take place in the Golgi apparatus?
There are two models for the operation of the Golgi apparatus in eukaryotic cells. As it is difficult to visualize the operation of cells at the molecular level in real time, scientists typically rely on static electron micrographs to see the morphology of organelles. As a result, the dynamic operation of these organelles can sometimes be unclear.
Cisternal Maturation Hypothesis
In the cisternal maturation hypothesis, the cisternae of the Golgi apparatus evolve. Proteins leave the endoplasmic reticulum, and enter the cis-Golgi. The cisterna of the cis-Golgi then matures, with its enzymatic contents and internal environment changing as it becomes the medial-Golgi, and, eventually, the trans-Golgi.
In this model, the proteins never physically leave their membrane-bound cisternae during their transit across the Golgi. Instead, the entire unit of contents remains within the evolving cisternae.
Vesicular Transport Hypothesis
In contrast to the cisternal maturation hypothesis, the vesicular transport hypothesis posits that the cis-, medial-, and trans-Golgi cisternae are more static structures. Instead of evolving around their contents, the contents are physically shuttled via vesicular intermediates from each cisterna to the next.
In the case of vesicular transport, vesicles are shuttled along microtubules. Motor proteins facilitate this movement, with unique proteins being used for each direction of movement along a microtubule.
Which of the following is most likely to take place in the Golgi apparatus?
Tap to reveal answer
Signal sequence removal and co-translational translocation are key events that occur in association with the endoplasmic reticulum, specifically the rough endoplasmic reticulum. The Golgi, in contrast, is specialized for the modification of polypeptides following their synthesis, such as through glycosylation.
Signal sequence removal and co-translational translocation are key events that occur in association with the endoplasmic reticulum, specifically the rough endoplasmic reticulum. The Golgi, in contrast, is specialized for the modification of polypeptides following their synthesis, such as through glycosylation.
← Didn't Know|Knew It →
There are two models for the operation of the Golgi apparatus in eukaryotic cells. As it is difficult to visualize the operation of cells at the molecular level in real time, scientists typically rely on static electron micrographs to see the morphology of organelles. As a result, the dynamic operation of these organelles can sometimes be unclear.
Cisternal Maturation Hypothesis
In the cisternal maturation hypothesis, the cisternae of the Golgi apparatus evolve. Proteins leave the endoplasmic reticulum, and enter the cis-Golgi. The cisterna of the cis-Golgi then matures, with its enzymatic contents and internal environment changing as it becomes the medial-Golgi, and, eventually, the trans-Golgi.
In this model, the proteins never physically leave their membrane-bound cisternae during their transit across the Golgi. Instead, the entire unit of contents remains within the evolving cisternae.
Vesicular Transport Hypothesis
In contrast to the cisternal maturation hypothesis, the vesicular transport hypothesis posits that the cis-, medial-, and trans-Golgi cisternae are more static structures. Instead of evolving around their contents, the contents are physically shuttled via vesicular intermediates from each cisterna to the next.
In the case of vesicular transport, vesicles are shuttled along microtubules. Motor proteins facilitate this movement, with unique proteins being used for each direction of movement along a microtubule.
A scientist is studying a set of proteins. She finds that protein A has a span of hydrophobic amino acids. Protein B also has a span of hydrophobic amino acids, but protein B also has carbohydrates added during modification in the Golgi, while protein A does not. If both protein A and protein B are hormone receptors, what role do the carbohydrate moieties most likely play?
There are two models for the operation of the Golgi apparatus in eukaryotic cells. As it is difficult to visualize the operation of cells at the molecular level in real time, scientists typically rely on static electron micrographs to see the morphology of organelles. As a result, the dynamic operation of these organelles can sometimes be unclear.
Cisternal Maturation Hypothesis
In the cisternal maturation hypothesis, the cisternae of the Golgi apparatus evolve. Proteins leave the endoplasmic reticulum, and enter the cis-Golgi. The cisterna of the cis-Golgi then matures, with its enzymatic contents and internal environment changing as it becomes the medial-Golgi, and, eventually, the trans-Golgi.
In this model, the proteins never physically leave their membrane-bound cisternae during their transit across the Golgi. Instead, the entire unit of contents remains within the evolving cisternae.
Vesicular Transport Hypothesis
In contrast to the cisternal maturation hypothesis, the vesicular transport hypothesis posits that the cis-, medial-, and trans-Golgi cisternae are more static structures. Instead of evolving around their contents, the contents are physically shuttled via vesicular intermediates from each cisterna to the next.
In the case of vesicular transport, vesicles are shuttled along microtubules. Motor proteins facilitate this movement, with unique proteins being used for each direction of movement along a microtubule.
A scientist is studying a set of proteins. She finds that protein A has a span of hydrophobic amino acids. Protein B also has a span of hydrophobic amino acids, but protein B also has carbohydrates added during modification in the Golgi, while protein A does not. If both protein A and protein B are hormone receptors, what role do the carbohydrate moieties most likely play?
Tap to reveal answer
The span of hydrophobic amino acids specified in the question suggests that these proteins are both transmembrane. Carbohydrate addition primarily occurs on portions of transmembrane proteins that face the extracellular environment, and serve to increase solubility in aqueous conditions. Out of the given options, only a modification to receptor specificity is an appropriate answer.
The span of hydrophobic amino acids specified in the question suggests that these proteins are both transmembrane. Carbohydrate addition primarily occurs on portions of transmembrane proteins that face the extracellular environment, and serve to increase solubility in aqueous conditions. Out of the given options, only a modification to receptor specificity is an appropriate answer.
← Didn't Know|Knew It →
There are two models for the operation of the Golgi apparatus in eukaryotic cells. As it is difficult to visualize the operation of cells at the molecular level in real time, scientists typically rely on static electron micrographs to see the morphology of organelles. As a result, the dynamic operation of these organelles can sometimes be unclear.
Cisternal Maturation Hypothesis
In the cisternal maturation hypothesis, the cisternae of the Golgi apparatus evolve. Proteins leave the endoplasmic reticulum, and enter the cis-Golgi. The cisterna of the cis-Golgi then matures, with its enzymatic contents and internal environment changing as it becomes the medial-Golgi, and, eventually, the trans-Golgi.
In this model, the proteins never physically leave their membrane-bound cisternae during their transit across the Golgi. Instead, the entire unit of contents remains within the evolving cisternae.
Vesicular Transport Hypothesis
In contrast to the cisternal maturation hypothesis, the vesicular transport hypothesis posits that the cis-, medial-, and trans-Golgi cisternae are more static structures. Instead of evolving around their contents, the contents are physically shuttled via vesicular intermediates from each cisterna to the next.
In the case of vesicular transport, vesicles are shuttled along microtubules. Motor proteins facilitate this movement, with unique proteins being used for each direction of movement along a microtubule.
In the cisternal maturation hypothesis, a change in which of the following is most likely to change the charge of the carboxyl and amino termini in a protein moving through the Golgi network?
There are two models for the operation of the Golgi apparatus in eukaryotic cells. As it is difficult to visualize the operation of cells at the molecular level in real time, scientists typically rely on static electron micrographs to see the morphology of organelles. As a result, the dynamic operation of these organelles can sometimes be unclear.
Cisternal Maturation Hypothesis
In the cisternal maturation hypothesis, the cisternae of the Golgi apparatus evolve. Proteins leave the endoplasmic reticulum, and enter the cis-Golgi. The cisterna of the cis-Golgi then matures, with its enzymatic contents and internal environment changing as it becomes the medial-Golgi, and, eventually, the trans-Golgi.
In this model, the proteins never physically leave their membrane-bound cisternae during their transit across the Golgi. Instead, the entire unit of contents remains within the evolving cisternae.
Vesicular Transport Hypothesis
In contrast to the cisternal maturation hypothesis, the vesicular transport hypothesis posits that the cis-, medial-, and trans-Golgi cisternae are more static structures. Instead of evolving around their contents, the contents are physically shuttled via vesicular intermediates from each cisterna to the next.
In the case of vesicular transport, vesicles are shuttled along microtubules. Motor proteins facilitate this movement, with unique proteins being used for each direction of movement along a microtubule.
In the cisternal maturation hypothesis, a change in which of the following is most likely to change the charge of the carboxyl and amino termini in a protein moving through the Golgi network?
Tap to reveal answer
The pH is the most direct condition that changes the charge found on the amino and carboxyl ends of a polypeptide. Protonation of both ends is favored by a low pH, while deprotonation is favored in more basic conditions.
The pH is the most direct condition that changes the charge found on the amino and carboxyl ends of a polypeptide. Protonation of both ends is favored by a low pH, while deprotonation is favored in more basic conditions.
← Didn't Know|Knew It →
There are two models for the operation of the Golgi apparatus in eukaryotic cells. As it is difficult to visualize the operation of cells at the molecular level in real time, scientists typically rely on static electron micrographs to see the morphology of organelles. As a result, the dynamic operation of these organelles can sometimes be unclear.
Cisternal Maturation Hypothesis
In the cisternal maturation hypothesis, the cisternae of the Golgi apparatus evolve. Proteins leave the endoplasmic reticulum, and enter the cis-Golgi. The cisterna of the cis-Golgi then matures, with its enzymatic contents and internal environment changing as it becomes the medial-Golgi, and, eventually, the trans-Golgi.
In this model, the proteins never physically leave their membrane-bound cisternae during their transit across the Golgi. Instead, the entire unit of contents remains within the evolving cisternae.
Vesicular Transport Hypothesis
In contrast to the cisternal maturation hypothesis, the vesicular transport hypothesis posits that the cis-, medial-, and trans-Golgi cisternae are more static structures. Instead of evolving around their contents, the contents are physically shuttled via vesicular intermediates from each cisterna to the next.
In the case of vesicular transport, vesicles are shuttled along microtubules. Motor proteins facilitate this movement, with unique proteins being used for each direction of movement along a microtubule.
A critical function of the Golgi apparatus is to modify proteins, typically by adding moieties and not by changing the actual amino acid sequence. A scientist finds that a large, negatively charged unit is added to a protein in the Golgi. Which of the following enzymes is most likely involved in this process?
There are two models for the operation of the Golgi apparatus in eukaryotic cells. As it is difficult to visualize the operation of cells at the molecular level in real time, scientists typically rely on static electron micrographs to see the morphology of organelles. As a result, the dynamic operation of these organelles can sometimes be unclear.
Cisternal Maturation Hypothesis
In the cisternal maturation hypothesis, the cisternae of the Golgi apparatus evolve. Proteins leave the endoplasmic reticulum, and enter the cis-Golgi. The cisterna of the cis-Golgi then matures, with its enzymatic contents and internal environment changing as it becomes the medial-Golgi, and, eventually, the trans-Golgi.
In this model, the proteins never physically leave their membrane-bound cisternae during their transit across the Golgi. Instead, the entire unit of contents remains within the evolving cisternae.
Vesicular Transport Hypothesis
In contrast to the cisternal maturation hypothesis, the vesicular transport hypothesis posits that the cis-, medial-, and trans-Golgi cisternae are more static structures. Instead of evolving around their contents, the contents are physically shuttled via vesicular intermediates from each cisterna to the next.
In the case of vesicular transport, vesicles are shuttled along microtubules. Motor proteins facilitate this movement, with unique proteins being used for each direction of movement along a microtubule.
A critical function of the Golgi apparatus is to modify proteins, typically by adding moieties and not by changing the actual amino acid sequence. A scientist finds that a large, negatively charged unit is added to a protein in the Golgi. Which of the following enzymes is most likely involved in this process?
Tap to reveal answer
The question notes that a large, negatively charges unit is added to the protein. This information, coupled with the answer options, tells us that the unit in question is a phosphate group. Phosphate groups are relatively large and carry a substantial negative charge. The addition of phosphate groups can help mediate protein activity by either activating or inactivating the molecule.
The question asks us to identify the enzyme responsible for adding the phosphate group to the protein; this function is unique to kinases.
The question notes that a large, negatively charges unit is added to the protein. This information, coupled with the answer options, tells us that the unit in question is a phosphate group. Phosphate groups are relatively large and carry a substantial negative charge. The addition of phosphate groups can help mediate protein activity by either activating or inactivating the molecule.
The question asks us to identify the enzyme responsible for adding the phosphate group to the protein; this function is unique to kinases.
← Didn't Know|Knew It →
One component of the immune system is the neutrophil, a professional phagocyte that consumes invading cells. The neutrophil is ferried to the site of infection via the blood as pre-neutrophils, or monocytes, ready to differentiate as needed to defend their host.
In order to leave the blood and migrate to the tissues, where infection is active, the monocyte undergoes a process called diapedesis. Diapedesis is a process of extravasation, where the monocyte leaves the circulation by moving in between endothelial cells, enters the tissue, and matures into a neutrophil.
Diapedesis is mediated by a class of proteins called selectins, present on the monocyte membrane and the endothelium. These selectins interact, attract the monocyte to the endothelium, and allow the monocytes to roll along the endothelium until they are able to complete diapedesis by leaving the vasculature and entering the tissues.
The image below shows monocytes moving in the blood vessel, "rolling" along the vessel wall, and eventually leaving the vessel to migrate to the site of infection.

Movement between cells, such as that carried out by monocytes in the passage, is typically blocked best by which kind of cell junction?
One component of the immune system is the neutrophil, a professional phagocyte that consumes invading cells. The neutrophil is ferried to the site of infection via the blood as pre-neutrophils, or monocytes, ready to differentiate as needed to defend their host.
In order to leave the blood and migrate to the tissues, where infection is active, the monocyte undergoes a process called diapedesis. Diapedesis is a process of extravasation, where the monocyte leaves the circulation by moving in between endothelial cells, enters the tissue, and matures into a neutrophil.
Diapedesis is mediated by a class of proteins called selectins, present on the monocyte membrane and the endothelium. These selectins interact, attract the monocyte to the endothelium, and allow the monocytes to roll along the endothelium until they are able to complete diapedesis by leaving the vasculature and entering the tissues.
The image below shows monocytes moving in the blood vessel, "rolling" along the vessel wall, and eventually leaving the vessel to migrate to the site of infection.

Movement between cells, such as that carried out by monocytes in the passage, is typically blocked best by which kind of cell junction?
Tap to reveal answer
Zona occludens block transport between cells by forming a zipper like boundary toward the apical surface of neighboring cells. These are also known as "tight junctions."
Zona adherens are also called adherens junctions or desmosones, and are designed to join cells together rather than to block transport between cells. Hemidesmosomes serve to link cells to an extracellular matrix or basement membrane, and gap junctions allow signal transduction between cells.
Zona occludens block transport between cells by forming a zipper like boundary toward the apical surface of neighboring cells. These are also known as "tight junctions."
Zona adherens are also called adherens junctions or desmosones, and are designed to join cells together rather than to block transport between cells. Hemidesmosomes serve to link cells to an extracellular matrix or basement membrane, and gap junctions allow signal transduction between cells.
← Didn't Know|Knew It →